Knowledge. Connectivity.
Experience. Delivery.
We are the leading provider of specialized services for the pharma, biotech and medtech industries.
PharmaLex Global Solutions
COVERING THE ENTIRE PRODUCT LIFECYCLE
Our solutions consist of core services, which are tailored to address each of your unique requirements. We also have extensive program management expertise, a key success factor within our service solutions.
Digital Innovation
We leverage artificial intelligence (AI) and machine learning automation, which is revolutionizing many industries through increased productivity, improved quality and enhancing return on investment.
Pharmaceutical
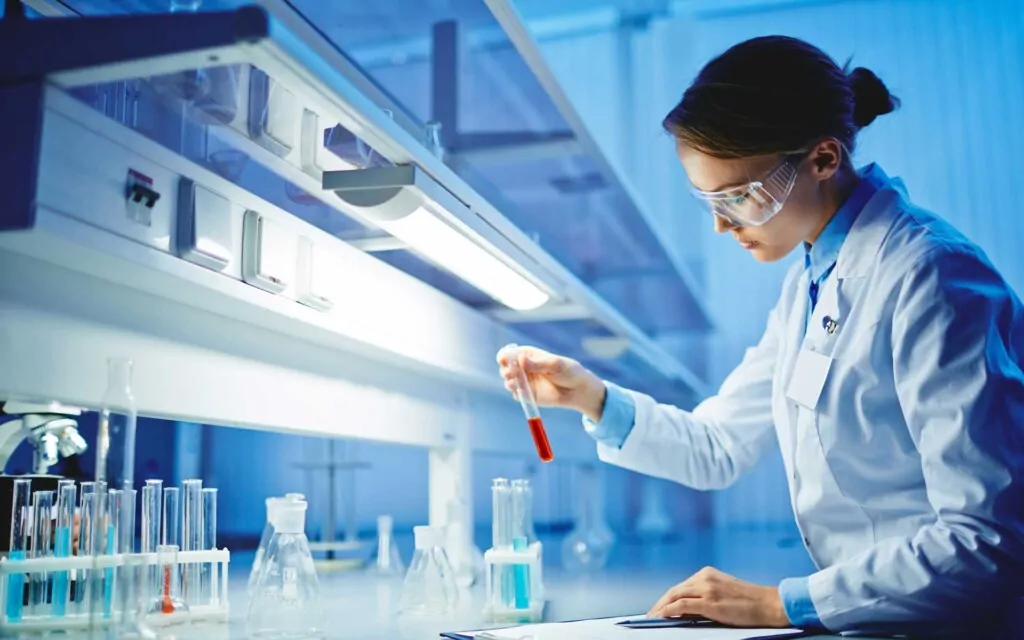

We offer global solutions covering the entire product lifecycle
Biotech
First-hand experience and a passion for science
MedTech
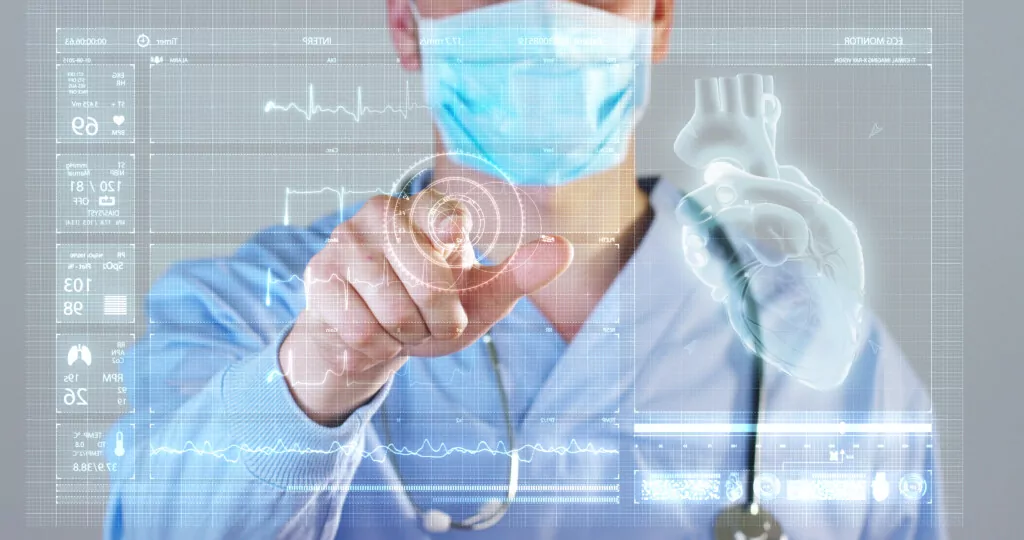

Proven methodology for long term success
PharmaLearn
LEARN FROM THE EXPERTS
High-quality, specialist training courses are enhancing our industry.
As Pharma consultants, we love to share our knowledge! Check out our blogs, webinars, articles, podcasts and more. Connect with our experts and stay up-to-date on all of the trends in the pharmaceutical, biotech and medical device industries.
- 18th April 2024
Corporate Social Responsibility
We are keenly focused on initiatives that revolve around education, empowerment, inclusivity and sustainability.
Global Contacts
We serve our clients through regional hubs, enabling personalized collaborate on projects at global, regional and local levels.
Europe
LATAM
MENA
North America
PharmaLex in Numbers
9 out of 10 top pharmaceutical companies are satisfied clients
Nationalities on staff, including former FDA and EMA experts
High satisfaction scores from satisfied clients
Customers include high percentage of small-medium-sized enterprises











